Post-Training Cosmos-H-Surgical-Simulator for Custom Surgical Robotics Dataset
Authors: Lukas Zbinden · Nigel Nelson · Maximilian Ofir Organization: NVIDIA
| Model | Workload | Use Case |
|---|---|---|
| Cosmos Predict 2.5 | Post-training | Surgical Robotics Simulation |
Motivation
Traditional surgical robot evaluation often requires expensive hardware and time-consuming physical setups, creating a significant bottleneck for rapid iteration and benchmarking. This recipe addresses this challenge by fine-tuning Cosmos Predict 2.5 to serve as a high-fidelity, action-conditioned surgical simulator. By predicting future video frames based on kinematics, the framework enables online evaluation and task success assessment within a purely digital environment. This work leverages the power of Cosmos to provide a safe, reproducible, and scalable pipeline that accelerates the development of autonomous surgical AI.
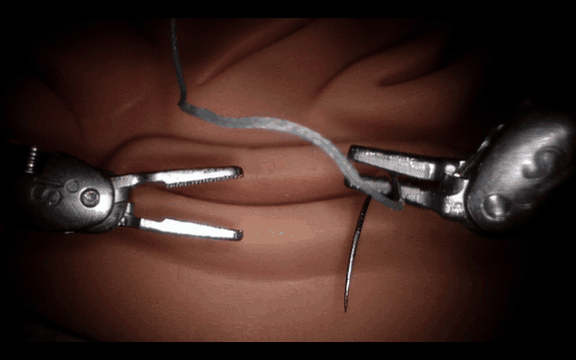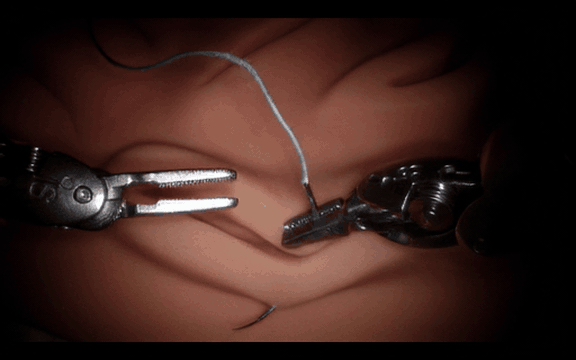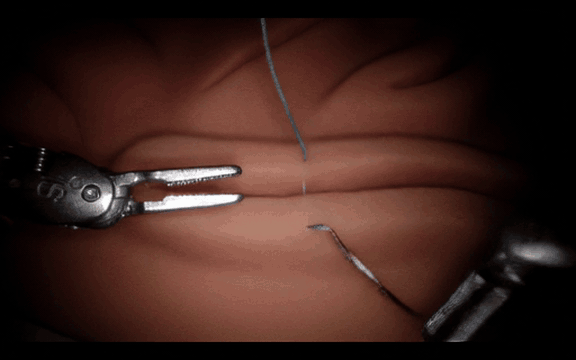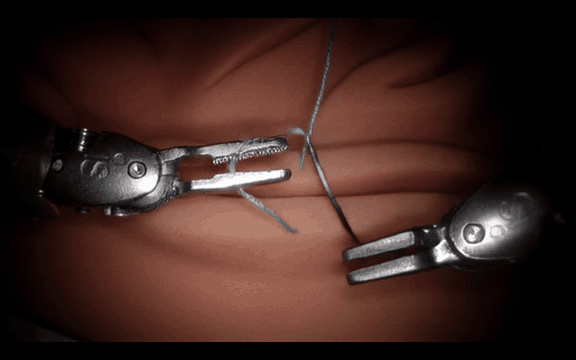
Post-Training Cosmos-H-Surgical-Simulator for Custom Surgical Robotics Dataset.
Overview
This tutorial guides you through post-training (finetuning) Cosmos-H-Surgical-Simulator, a version of Cosmos Predict 2.5 pre-trained on the Open-H embodiment surgical robotics datasets, on the downstream SutureBot dataset. The resulting model functions as a learned simulator for policy evaluation and synthetic data generation, implicitly capturing both robot kinematics and task-relevant environment dynamics.
The approach builds on Cosmos-Surg-dVRK and uses the public SutureBot dataset, which contains endoscopic video paired with kinematic action sequences from the da Vinci Research Kit (dVRK), as a custom surgical dataset for downstream finetuning. While demonstrated on surgical robotics, this tutorial generalizes to other robotic systems and broader embodied AI applications.
Cosmos-H-Surgical-Simulator was finetuned on the Open-H embodiment surgical datasets with a unified 44-dimensional action space, where the CMR Surgical Versius uses the full 44D (30D actions + 14D state conditioning) and every other embodiment (dVRK JHU, Stanford, Hamlyn, etc.) is zero-padded to 44D. For the SutureBot downstream finetuning described in this tutorial, SutureBot's native 20D actions are zero-padded to 44D to remain compatible with the pre-trained model's action MLP. The 24 trailing zeros occupy the same positions as CMR's extra channels, which the Cosmos-predict2.5 model has already learned can be zero since all non-CMR Open-H datasets had zeros there during pre-finetuning.
Because the Cosmos-H-Surgical-Simulator has already learned surgical visual appearance, dVRK kinematics, and action-conditioned video dynamics from the diverse Open-H embodiment collection, which itself includes closely related dVRK suturing data, downstream finetuning on a new surgical robotics dataset like SutureBot is expected to converge in substantially fewer iterations than training from the base Cosmos-predict2.5 model alone.
Table of Contents
- Prerequisites
- Preparing Data
- Bring Your Own Dataset
- Action Format
- Model Configuration
- Finetuning
- Inference and Evaluation
- Results
- Downloading Artifacts
- Further Reading
1. Prerequisites
Recommended setup: Build the Docker image (step 1.5) and run all finetuning and inference commands inside the container. Docker handles all CUDA, PyTorch, and Cosmos dependencies without any host-level configuration. The host Python environment (step 1.4) is only needed for the data preparation scripts in step 2.
Complete the steps below in order.
1.1 Clone the Cosmos Cookbook
Clone the Cosmos Cookbook repository, which contains the data preparation scripts and documentation for this tutorial:
git clone https://github.com/nvidia-cosmos/cosmos-cookbook.git
cd cosmos-cookbook/docs/recipes/post_training/predict2_5/surgical_robotics
1.2 System Setup (Fresh Cloud Instances)
If you are working on a freshly provisioned cloud instance (e.g. via brev or similar), run the system setup script first. It configures Docker and containerd to use the largest available drive, cleans up logs to free disk space, and sets DNS for Docker:
Skip this step if your instance already has Docker configured with sufficient storage.
1.3 Clone the Cosmos-H-Surgical-Simulator Repository
Clone the Cosmos-H-Surgical-Simulator repository (a fork of cosmos-predict2.5 with this tutorial's code changes applied):
git clone https://github.com/NVIDIA-Medtech/Cosmos-H-Surgical-Simulator.git
cd Cosmos-H-Surgical-Simulator
1.4 Run the Setup Guide
Follow the Setup guide: install system dependencies, uv, Python env (uv sync --extra=cu128), and HF CLI. Finish all steps before continuing.
Important: Run
uv syncas the same user who will run the data preparation scripts (step 2), not as root. Ifuv syncwas run as root, remove and recreate the venv as the current user:
1.5 Build the Docker Image (for Containerized Runs)
If you will run finetuning via Docker (recommended), build the image from the Cosmos-H-Surgical-Simulator repository:
cd /path/to/Cosmos-H-Surgical-Simulator
docker build -f Dockerfile -t cosmos-predict2.5:local .
export COSMOS_CONTAINER_IMAGE=cosmos-predict2.5:local
1.6 Configure Environment Variables
Navigate back to the cookbook recipe directory (cloned in step 1.1). All data preparation scripts, the finetuning launcher, and the environment template live here:
All paths and credentials are managed through a single environment file. Copy the template, fill in every value, then source it before running any command in this tutorial:
cp scripts/env.sh.template scripts/env.sh
# Edit scripts/env.sh — fill in all values (see descriptions below)
source scripts/env.sh
scripts/env.shis gitignored and must never be committed — it contains your API keys and machine-specific paths.
The variables and why they matter:
| Variable | Description |
|---|---|
HF_HOME |
HuggingFace cache for model weights and LeRobot datasets. Needs ~100 GB. Authenticate first: hf auth login |
IMAGINAIRE_OUTPUT_ROOT |
Root directory for training checkpoints (saved every 200 steps). Needs ~500 GB for a full run. |
WANDB_API_KEY |
Weights & Biases API key for experiment tracking. Get yours at wandb.ai/settings. Required to monitor training loss and convergence. |
COSMOS_CONTAINER_IMAGE |
Docker image tag built in step 1.5 (default: cosmos-predict2.5:local). |
COSMOS_CODE_PATH |
Absolute path to the Cosmos-H-Surgical-Simulator repo cloned in step 1.3. |
SUTUREBOT_LEROBOT_PATH |
Absolute path to the converted LeRobot dataset (step 2.3). Start with the mini dataset path; swap for the full dataset when ready. |
COSMOS_H_CKPT_PATH |
Path to the downloaded Cosmos-H DCP checkpoint directory (iter_000023000/). |
SAVE_ROOT |
Output directory for inference videos (step 5.3). |
2. Preparing Data
All training data must be in LeRobot v3 format — a standardized structure used by the Cosmos-H training pipeline. This section converts the public SutureBot dataset to that format. To adapt the workflow to a different robot or task, see step 2.4.
Getting started: Use the mini dataset (step 2.3) to verify the full pipeline end-to-end before committing to a full dataset conversion or long training run. Prerequisites: The scripts in this section require the packages installed in step 1.4. Activate the venv before running any commands:
2.1 About the SutureBot Dataset
SutureBot is a dataset for autonomous end-to-end suturing on the dVRK, covering subtasks like needle pickup, needle insertion, and knot tying. It provides multi-camera surgical video paired with robot kinematics to support imitation learning and evaluation of VLA/robotic policies. SutureBot contains about 1,890 demonstrations, amounting to 6 hours of video or 629,183 samples.
| Needle pickup | Needle insertion | Knot tying |
|---|---|---|
2.2 Download
Set the dataset destination and run the download script:
Unpack zip files:
2.3 Convert to LeRobot Dataset Format
To be compatible with Cosmos data processing, convert the raw SutureBot data to the LeRobot Dataset format.
The converted dataset is written to $HF_HOME/lerobot/<repo_id> by default (lerobot follows the HuggingFace cache convention). Since HF_HOME is already set from step 1.6, no extra path configuration is needed. To override the output location, set HF_LEROBOT_HOME before running.
Mini dataset (for quick testing): because full conversion takes about 1.5–2.5 hours, you can create a small subset first:
python3 -u scripts/create_mini_suturebot.py \
--source $SUTUREBOT_DATASET_DIR \
--output $SUTUREBOT_DATASET_DIR/SutureBot_mini \
--max-episodes 3 \
--tissue tissue_1
This copies a subset of episodes and then runs convert_suturebot_to_lerobot_v3.py on that folder. Add --no-convert to only create the mini folder. The LeRobot dataset is written to $HF_HOME/lerobot/suturebot_lerobot_mini.
Full dataset conversion: run the converter directly on the full dataset (lerobot==0.3.3 is expected):
The output is written to $HF_HOME/lerobot/suturebot_lerobot.
2.4 Bring Your Own Dataset
To fine-tune on a custom robot or task, your data must be in LeRobot v3 format:
<dataset_root>/
├── meta/
│ ├── info.json # fps, feature names, episode/frame counts
│ ├── episodes.json
│ └── stats.json # action mean/std for normalization
├── data/chunk-000/
│ └── episode_000000.parquet # per-frame observations and actions
└── videos/chunk-000/<camera_key>/
└── episode_000000.mp4
The LeRobot library provides utilities for building and validating datasets in this format. Once your dataset is ready, three additional steps integrate it with the training pipeline:
- Register an embodiment tag — add a new entry to
EmbodimentTagand a config block ingroot_configs.py(steps 3.1 and 3.4). - Register the dataset — add train/val dataset entries in
data.pyand an experiment config (steps 3.2–3.3). - Update inference — change
embodiment="suturebot"ininference_dvrk.pyto your new embodiment tag.
See Model Configuration for a concrete example of all three changes applied for SutureBot.
2.5 Action Format
Understanding the action representation is important for interpreting inference results and for adapting this workflow to other robots.
Dataset (SutureBot LeRobot)
Each frame in the converted parquet files stores a 20-dimensional absolute Cartesian setpoint for the two PSM arms:
| Dimensions | Field | Description |
|---|---|---|
| 0–2 | psm1_xyz |
PSM1 end-effector position (metres) |
| 3–8 | psm1_rot6d |
PSM1 orientation as first two rows of rotation matrix |
| 9 | psm1_jaw |
PSM1 jaw angle (radians) |
| 10–12 | psm2_xyz |
PSM2 end-effector position (metres) |
| 13–18 | psm2_rot6d |
PSM2 orientation as first two rows of rotation matrix |
| 19 | psm2_jaw |
PSM2 jaw angle (radians) |
At training and inference time, RelativeActionTransform converts each 13-frame chunk into 20D per-chunk relative actions:
- Translation: global frame delta —
Δxyz = xyz_target − xyz_base - Rotation: local frame delta —
ΔR = R_base.T @ R_targetin 6D form (first two rows of the relative rotation matrix) - Jaw: absolute setpoint (not a delta)
The base pose is always the first frame of the chunk. Normalization uses stats.json computed from the SutureBot dataset itself (mean/std of per-chunk deltas).
Pre-trained Cosmos-H Model
The Cosmos-H-Surgical-Simulator checkpoint was pre-trained on the Open-H community surgical dataset (~3M frames across 9 institutions and 11 robot types). It uses a 44-dimensional unified action space: each robot contributes its native action dimensions, with trailing zeros padding to 44D.
SutureBot-type data (suturebot_2, suturebot_3, suturebot_tissue_2 from JHU) was included in pre-training under the jhu_dvrk_mono embodiment, processed with GenericRelativeActionTransform (per-key relative xyz + rot6d) and normalized with Open-H community statistics (stats_cosmos.json).
Fine-tuning and Inference Alignment
Fine-tuning registers SutureBot as a distinct embodiment (suturebot) that uses RelativeActionTransform with the dataset's own stats.json. The inference script zero-pads actions from 20D to 44D before passing them to the model, matching the padding applied during fine-tuning.
Note: Running inference with the pre-trained Cosmos-H checkpoint on SutureBot data will produce near-static output. The pre-trained model's action embedder was calibrated to Open-H statistics, while the SutureBot dataset uses a different normalization distribution, causing the action signal to be misinterpreted. Meaningful motion generation requires fine-tuning on the SutureBot dataset first (step 4).
3. Model Configuration
The finetuning is performed at 288x512 resolution (to match the Cosmos-H-Surgical-Simulator pre-finetuning) with a 12-frame prediction horizon.
If you cloned the Cosmos-H-Surgical-Simulator repository in step 1.3, these code changes are already applied. Skip this section and go to Finetuning.
If you are using the upstream cosmos-predict2.5 repository instead (v1.4.1), you must apply the following changes. The subsections below document them for reference.
3.1 Register the 'suturebot' embodiment
File: cosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/data/embodiment_tags.py
Add a new SUTUREBOT = "suturebot" entry to the EmbodimentTag enum.
3.2 Configure the 2B model for SutureBot
File: cosmos_predict2/_src/predict2/action/configs/action_conditioned/experiment/exp_2B_action_conditioned_rectify_flow_gr00t.py
Add a new AC_CHUNK_SINGLE_VIEW_2B_SUTUREBOT_13FRAME_4NODES_OSS config dict defining: single-view SutureBot dataset references, action dimension of 20, batch size of 4, learning rate 4e-5, and weight decay 0.1.
3.3 Define SutureBot data loading
File: cosmos_predict2/_src/predict2/action/configs/action_conditioned/data.py
Register suturebot_train and suturebot_val datasets and dataloaders using the LeRobotDataset class with 13 frames, embodiment="suturebot", and max_pixels=1920*1080.
3.4 Add SutureBot configuration (resolution, delta actions, normalization)
File: cosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/groot_configs.py
Add suturebot embodiment config with timestep_interval=3, resolution 960x720, and switch normalization to mean_std. Add RelativeActionTransform to the transform pipeline.
3.5–3.6 Bugfixes in the Cosmos OSS code
Files:
cosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/data/transform/video.pycosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/data/transform/concat.py
Fix .split(".") calls to .split(".", 1) to handle keys with multiple dots (e.g. video.observation.images.main).
3.7 Relative action computation
File: cosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/data/transform/state_action.py
Add RelativeActionTransform class and helper functions compute_rel_actions / compute_rel_actions_local that compute kinematic delta actions following Stanford's UMI implementation.
3.8 Video loading bugfix (AV1 codec)
File: cosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/utils/video.py
Fix frame matching logic by using closest-timestamp matching instead of sequential loading, which broke for certain video codecs (AV1).
3.9 Dataset class changes for delta actions
File: cosmos_predict2/_src/predict2/action/datasets/gr00t_dreams/data/dataset.py
Simplify the LeRobotDataset to use the RelativeActionTransform in the pipeline instead of manually computing delta actions in the __getitem__ method.
4. Finetuning
Fine-tuning adapts the Cosmos-H-Surgical-Simulator checkpoint to your dataset by jointly training the action embedder and diffusion backbone.
Prerequisites
Before starting, ensure you have completed:
- Build the Docker Image — Docker image built (
cosmos-predict2.5:local) - Configure Environment Variables —
scripts/env.shfilled in and sourced (source scripts/env.sh) - Convert to LeRobot Dataset Format — LeRobot dataset prepared (start with the mini dataset)
Download Cosmos-H-Surgical-Simulator Checkpoint
Download the pre-trained DCP checkpoint from HuggingFace:
hf download nvidia/Cosmos-H-Surgical-Simulator \
--include "checkpoints/iter_000023000/model/*" \
--local-dir /path/to/checkpoints
export COSMOS_H_CKPT_PATH=/path/to/checkpoints/checkpoints/iter_000023000
COSMOS_H_CKPT_PATH points to the iter_000023000 directory (not model/ — the trainer appends that internally). If left unset, training warm-starts from the base Cosmos 2B model.
Run Finetuning
Using Docker (recommended):
export COSMOS_CODE_PATH=/path/to/Cosmos-H-Surgical-Simulator
export SUTUREBOT_LEROBOT_PATH=/path/to/suturebot_lerobot_mini # mini dataset (step 2.3); swap for full dataset when ready
export COSMOS_H_CKPT_PATH=/path/to/checkpoints/checkpoints/iter_000023000
export IMAGINAIRE_OUTPUT_ROOT=/path/to/training_output
export COSMOS_CONTAINER_IMAGE=cosmos-predict2.5:local
export HF_HOME=/path/to/huggingface_cache
./scripts/run_finetuning_standalone.sh
Set NGPUS=<n> to control GPU count (default: all available). Set WANDB_API_KEY=<key> to enable W&B logging.
Reference run (8×H100, mini dataset): the exact configuration used in this tutorial:
export COSMOS_CODE_PATH=/ephemeral/Cosmos-H-Surgical-Simulator
export SUTUREBOT_LEROBOT_PATH=/ephemeral/data/suturebot_lerobot_mini
export COSMOS_H_CKPT_PATH=/ephemeral/checkpoints/checkpoints/iter_000023000
export IMAGINAIRE_OUTPUT_ROOT=/ephemeral/checkpoints/training_output
export COSMOS_CONTAINER_IMAGE=cosmos-predict2.5:local
export HF_HOME=/ephemeral/cache/huggingface
export WANDB_API_KEY=your_api_key_here # optional
./scripts/run_finetuning_standalone.sh
Without Docker (host venv):
export SUTUREBOT_LEROBOT_PATH=/path/to/suturebot_lerobot
export COSMOS_CODE_PATH=/path/to/Cosmos-H-Surgical-Simulator
export COSMOS_H_CKPT_PATH=/path/to/checkpoints/checkpoints/iter_000023000
export IMAGINAIRE_OUTPUT_ROOT=/path/to/training_output
./scripts/run_finetuning_standalone.sh
Key Training Parameters
| Parameter | Default | How to Change |
|---|---|---|
| GPUs | all available | NGPUS=<n> env var |
| Batch size (per GPU) | 4 (global: NGPUS × 4) |
Edit experiment config in step 3.2 |
| Learning rate | 4e-5 | Append optimizer.lr=<value> to the training command |
| Checkpoint save interval | every 200 steps | Change checkpoint.save_iter=200 in run_finetuning_standalone.sh |
| W&B logging | off | Set WANDB_API_KEY |
Training Details
Checkpoints are saved every 200 steps to:
${IMAGINAIRE_OUTPUT_ROOT}/cosmos_predict2_action_conditioned/official_runs_vid2vid/cosmos_predict2p5_2B_action_conditioned_suturebot_13frame_4nodes_release_oss/checkpoints/
Expected training time on 8×H100 PCIe (~20 steps/min):
| Steps | Time | Notes |
|---|---|---|
| 5,000 | ~4 h | Early improvement visible on mini dataset |
| 13,000 | ~11 h | Reference run used in this tutorial |
| 23,000 | ~20 h | Published Cosmos-H-Surgical-Simulator checkpoint |
For the mini dataset (3 episodes), the model converges quickly — suitable for pipeline verification. For the full SutureBot dataset (~1,890 episodes), plan for 15,000–23,000 steps for strong results. Training time scales approximately linearly with fewer GPUs (e.g., 1×H100 ≈ 8× longer).
Note: The step counts above were established with an intermediate Cosmos-H-Surgical-Simulator checkpoint (pre-finetuned on Open-H). Convergence using the final checkpoint is expected to be substantially faster since the model already encodes surgical visual priors and dVRK action dynamics. This may be the case as well for any downstream surgical robotics dataset. Monitor validation loss and sample quality to determine an appropriate early stopping point.
To use a fine-tuned checkpoint for inference, convert the DCP to a .pt file (see step 5.1).
5. Inference and Evaluation
This tutorial is grounded in the methodology of Cosmos-Surg-dVRK, which validated the world model by comparing policy success rates in Cosmos simulation against real-world robot execution (Pearson r = 0.718, p < 0.001).
The inference_dvrk.py script runs autoregressive video generation for policy evaluation:
- Loads only the first frame from the dataset as initial conditioning
- Generates frames using ground-truth actions from the dataset
- Uses each chunk's last predicted frame as conditioning for the next chunk
- Stitches all chunks into a full episode video
5.1 Convert Checkpoint
Training produces distributed checkpoints (DCP) that must be converted to a single .pt file before inference. The conversion script lives inside the Cosmos-H-Surgical-Simulator repo.
Set COSMOS_CODE_PATH, CHECKPOINTS_DIR, and CHECKPOINT_ITER for whichever checkpoint you want to convert:
Pre-trained checkpoint (downloaded from HuggingFace in step 4):
# Download only the model weights (skip optimizer/scheduler for inference)
hf download nvidia/Cosmos-H-Surgical-Simulator \
--include "checkpoints/iter_000023000/model/*" \
--local-dir /path/to/checkpoints
COSMOS_CODE_PATH=/path/to/Cosmos-H-Surgical-Simulator
CHECKPOINTS_DIR=/path/to/checkpoints/checkpoints
CHECKPOINT_ITER=iter_000023000
Fine-tuned checkpoint (from your training run in step 4):
COSMOS_CODE_PATH=/path/to/Cosmos-H-Surgical-Simulator
CHECKPOINTS_DIR=$IMAGINAIRE_OUTPUT_ROOT/cosmos_predict2_action_conditioned/official_runs_vid2vid/cosmos_predict2p5_2B_action_conditioned_suturebot_13frame_4nodes_release_oss/checkpoints
CHECKPOINT_ITER=iter_000013000 # replace with your chosen iteration
Once those variables are set, run the conversion:
Using Docker (recommended):
docker run --rm \
-v $COSMOS_CODE_PATH:/workspace \
-v $CHECKPOINTS_DIR:$CHECKPOINTS_DIR \
-w /workspace \
$COSMOS_CONTAINER_IMAGE \
bash -c "source .venv/bin/activate 2>/dev/null || true && \
python scripts/convert_distcp_to_pt.py \
$CHECKPOINTS_DIR/$CHECKPOINT_ITER/model \
$CHECKPOINTS_DIR/$CHECKPOINT_ITER"
Without Docker (host venv):
cd $COSMOS_CODE_PATH
source .venv/bin/activate
python scripts/convert_distcp_to_pt.py \
$CHECKPOINTS_DIR/$CHECKPOINT_ITER/model \
$CHECKPOINTS_DIR/$CHECKPOINT_ITER
This creates three files in $CHECKPOINTS_DIR/$CHECKPOINT_ITER/:
model.pt— full checkpoint (regular + EMA weights)model_ema_fp32.pt— EMA weights in float32model_ema_bf16.pt— EMA weights in bfloat16 (recommended for inference)
5.2 Copy Inference Script
From the cookbook recipe directory, copy the inference script into the Cosmos-H-Surgical-Simulator repo:
5.3 Run Inference
Set the paths (reuse variables from step 5.1, or redefine them here for a new terminal session):
COSMOS_CODE_PATH=/path/to/Cosmos-H-Surgical-Simulator
CHECKPOINTS_DIR=/path/to/checkpoints/checkpoints # same value as in step 5.1
CHECKPOINT_ITER=iter_000023000 # whichever iter you converted
SUTUREBOT_LEROBOT_PATH=$HF_HOME/lerobot/suturebot_lerobot_mini # mini dataset (step 2.3)
# SUTUREBOT_LEROBOT_PATH=$HF_HOME/lerobot/suturebot_lerobot # full dataset
SAVE_ROOT=/path/to/results/dvrk_eval
The script generates rollouts given ground-truth kinematic action trajectories and an initial frame from the dataset.
Note on
--experiment: The inference command uses the samesuturebotexperiment config (cosmos_predict2p5_2B_action_conditioned_suturebot_13frame_4nodes_release_oss) as training. The inference pipeline reads this config to set up the model architecture, while data loading is handled separately ininference_dvrk.pyusingembodiment="suturebot"and the dataset's ownstats.json.
Using Docker (recommended):
docker run --rm --gpus all \
-v $COSMOS_CODE_PATH:/workspace \
-v $CHECKPOINTS_DIR:$CHECKPOINTS_DIR \
-v $SUTUREBOT_LEROBOT_PATH:$SUTUREBOT_LEROBOT_PATH \
-v $SAVE_ROOT:$SAVE_ROOT \
-w /workspace \
$COSMOS_CONTAINER_IMAGE \
bash -c "source .venv/bin/activate 2>/dev/null || true && \
CUDA_VISIBLE_DEVICES=0 PYTHONPATH=. python \
cosmos_predict2/_src/predict2/action/inference/inference_dvrk.py \
--experiment=cosmos_predict2p5_2B_action_conditioned_suturebot_13frame_4nodes_release_oss \
--ckpt_path $CHECKPOINTS_DIR/$CHECKPOINT_ITER/model_ema_bf16.pt \
--dataset_path $SUTUREBOT_LEROBOT_PATH \
--save_root $SAVE_ROOT \
--data_split train \
--episode_ids 0,1,2 \
--save_comparison"
Without Docker (host venv):
cd $COSMOS_CODE_PATH
source .venv/bin/activate
CUDA_VISIBLE_DEVICES=0 PYTHONPATH=. python cosmos_predict2/_src/predict2/action/inference/inference_dvrk.py \
--experiment=cosmos_predict2p5_2B_action_conditioned_suturebot_13frame_4nodes_release_oss \
--ckpt_path $CHECKPOINTS_DIR/$CHECKPOINT_ITER/model_ema_bf16.pt \
--dataset_path $SUTUREBOT_LEROBOT_PATH \
--save_root $SAVE_ROOT \
--data_split train \
--episode_ids 0,1,2 \
--save_comparison
Note: The
--data_split trainflag is used here because the mini dataset from step 2.3 contains only atrainsplit. For a full dataset conversion (which producestrain/testsplits), use--data_split test.
The --save_comparison flag generates side-by-side videos (ground truth on the left, predicted on the right).
5.4 Inference Results
The following metrics are from the reference run (iter_000013000, mini dataset, 1×H100 PCIe):
| Metric | Value |
|---|---|
| GPU memory | ~20 GB |
| Denoising speed | ~9.9 it/s (36 steps/chunk) |
| Time per 12-frame chunk | ~4 s |
| Time per episode (~10 chunks) | ~40–45 s |
Output files are written to $SAVE_ROOT:
dvrk_eval/
├── predicted/
│ ├── episode_0000.mp4 # generated video
│ ├── episode_0001.mp4
│ └── episode_0002.mp4
├── comparison/
│ ├── episode_0000.mp4 # side-by-side: ground truth (left) vs predicted (right)
│ ├── episode_0001.mp4
│ └── episode_0002.mp4
└── action_log.json # logged action data per episode
Base model vs fine-tuned: Running inference with the pre-trained Cosmos-H checkpoint (without SutureBot fine-tuning) produces near-static output — the predicted frames barely change from the conditioning frame. This is expected: the pre-trained model uses Open-H action statistics, which are incompatible with the SutureBot normalization (see step 2.5). Fine-tuning on SutureBot data (step 4) is required to generate meaningful motion.
5.5 Swapping in a Surgical Policy
To evaluate a surgical policy (a VLA model) instead of ground-truth actions, modify the inference loop in inference_dvrk.py:
# Current (GT actions from dataset):
actions = data["action"].numpy()
# With a policy:
actions = policy.predict(current_frame) # Returns (12, action_dim)
The finetuned Cosmos model expects normalized action sequences matching the shape (chunk_size, action_dim) and following the relative action formulation used during training.
Note: Running Cosmos with a policy's output actions generates video rollouts (MP4 files) for manual review. To automate this evaluation process, Cosmos-Reason2 can be post-trained to serve as a judge, automatically detecting task successes, failures, and physics anomalies.
6. Results
The post-trained Cosmos-H-Surgical-Simulator model generates faithful and highly realistic rollouts compared to the ground-truth video. Below are side-by-side comparison videos (ground truth on the left, predicted on the right) from the reference run. Run inference as described in step 5.3 to generate these videos.
| Task | Ground Truth | Post-Trained Model |
|---|---|---|
| Pickup & Handover | ||
| Throw & Extraction | ||
| Knot Tie |
7. Downloading Artifacts
After running the tutorial on a cloud instance (e.g. brev), use the commands below to pull results to your local machine. Replace <instance-name> with your brev instance name (visible in brev ls).
Each artifact has two download options:
brev copy— purpose-built for brev instances; no SSH config requiredrsync— works with any SSH-accessible host; brev adds instance entries to~/.ssh/configso<instance-name>works directly as a hostname
Evaluation Videos
Side-by-side comparison videos and predicted rollouts from step 5.3:
# brev
brev copy <instance-name>:/ephemeral/results/dvrk_eval/finetuned_iter13000_mini10ep dvrk_eval_iter13000
# rsync
rsync -avz --progress <instance-name>:/ephemeral/results/dvrk_eval/finetuned_iter13000_mini10ep/ dvrk_eval_iter13000/
Converted Model Checkpoint
The EMA bf16 checkpoint (4 GB) produced by step 5.1 — suitable for inference and further fine-tuning:
# brev
brev copy <instance-name>:/ephemeral/checkpoints/converted_iter13000/model_ema_bf16.pt model_ema_bf16.pt
# rsync
rsync -avz --progress <instance-name>:/ephemeral/checkpoints/converted_iter13000/model_ema_bf16.pt ./model_ema_bf16.pt
To download the full checkpoint directory (includes fp32 and full weights, ~24 GB total):
# brev
brev copy <instance-name>:/ephemeral/checkpoints/converted_iter13000 converted_iter13000
# rsync
rsync -avz --progress <instance-name>:/ephemeral/checkpoints/converted_iter13000/ converted_iter13000/
LeRobot Dataset
The converted mini dataset from step 2.3 (~few GB depending on episode count):
# brev
brev copy <instance-name>:/ephemeral/cache/huggingface/lerobot/suturebot_lerobot_mini suturebot_lerobot_mini
# rsync
rsync -avz --progress <instance-name>:/ephemeral/cache/huggingface/lerobot/suturebot_lerobot_mini/ suturebot_lerobot_mini/
Further Reading
- Cosmos-H-Surgical-Simulator repo — Cosmos-predict2.5 fine-tuned on the Open-H embodiment dataset
- Cosmos-H-Surgical-Simulator checkpoint - Cosmos-H-Surgical-Simulator checkpoint on Hugging Face.
- Open-H embodiment — Open-H-Embodiment community‑driven dataset
- Cosmos Predict 2.5 — Model weights and documentation
- SutureBot — A Precision Framework & Benchmark for Autonomous End-to-End Suturing
- Cosmos-Surg-dVRK — World foundation model-based automated online evaluation of surgical robot policy learning
- The da Vinci Research Kit — A community effort supporting research in telerobotic surgery
Document Information
Publication Date: March 15, 2026
Citation
If you use this recipe or reference this work, please cite it as:
@misc{cosmos_cookbook_surgical_robotics_2026,
title={Post-Training Cosmos-H-Surgical-Simulator for Custom Surgical Robotics Dataset},
author={Zbinden, Lukas and Nelson, Nigel and Ofir, Maximilian},
year={2026},
month={March},
howpublished={\url{https://nvidia-cosmos.github.io/cosmos-cookbook/recipes/post_training/predict2_5/surgical_robotics/post_training.html}},
note={NVIDIA Cosmos Cookbook}
}
Suggested text citation:
Lukas Zbinden, Nigel Nelson & Maximilian Ofir (2026). Post-Training Cosmos-H-Surgical-Simulator for Custom Surgical Robotics Dataset. In NVIDIA Cosmos Cookbook. Accessible at https://nvidia-cosmos.github.io/cosmos-cookbook/recipes/post_training/predict2_5/surgical_robotics/post_training.html